Welcome to the home of 'Tissue sampling'
These pages include chapters on sampling, fixation and processing of tissue through to staining, microscopy and whole slide imaging. Atlases of normal tissues and special stains can be found at the end of the chapter on 'Staining'. The books 'Tissue processing: From patient to pathologist' and 'Pathological bodies' are both available on Amazon.
In addition to 'Pathological bodies', you can click on the 'PathoFocus' tab above to access my other website on the pathology of disease processes. The 'Publications' tab will take you to my books, journal and web-based publications and blogs and international presentations. I do hope you enjoy your visit and please feel free to post a comment.Following your visit, why not pop across to the 'Atlas of medical foreign bodies'. Compiled by Dr Yale Rosen, a New York pathologist, you can access it at https://t.co/tJ8RajYSlL
These pages include chapters on sampling, fixation and processing of tissue through to staining, microscopy and whole slide imaging. Atlases of normal tissues and special stains can be found at the end of the chapter on 'Staining'. The books 'Tissue processing: From patient to pathologist' and 'Pathological bodies' are both available on Amazon.
In addition to 'Pathological bodies', you can click on the 'PathoFocus' tab above to access my other website on the pathology of disease processes. The 'Publications' tab will take you to my books, journal and web-based publications and blogs and international presentations. I do hope you enjoy your visit and please feel free to post a comment.Following your visit, why not pop across to the 'Atlas of medical foreign bodies'. Compiled by Dr Yale Rosen, a New York pathologist, you can access it at https://t.co/tJ8RajYSlL
Sampling of tissue outside the laboratory
Biopsies are invasive procedures and should be performed as a result of clinical indications and with a genuine possibility of therapeutic benefit. For the patient, tissue sampling is not without risk and the most important of them are described at the end of this chapter. Imaging methods are routinely used for obtaining certain tissue samples although most radiographic evaluations are confined to breast disease. Microcalcification can sometime mark the site of a neoplastic lesion in the breast where radiology can be used to locate tiny lesions that may not be obvious on palpation or laboratory examination. Routine breast radiography should certainly be done when initial sampling fails to reveal a lesion that has been picked up on mammography. There are several imaging methods that can be employed in the sampling of breast and other tissues and these are included in the alphabetic list of sampling methods below.
Biopsies are invasive procedures and should be performed as a result of clinical indications and with a genuine possibility of therapeutic benefit. For the patient, tissue sampling is not without risk and the most important of them are described at the end of this chapter. Imaging methods are routinely used for obtaining certain tissue samples although most radiographic evaluations are confined to breast disease. Microcalcification can sometime mark the site of a neoplastic lesion in the breast where radiology can be used to locate tiny lesions that may not be obvious on palpation or laboratory examination. Routine breast radiography should certainly be done when initial sampling fails to reveal a lesion that has been picked up on mammography. There are several imaging methods that can be employed in the sampling of breast and other tissues and these are included in the alphabetic list of sampling methods below.
Advanced Breast Biopsy Instrumentation (ABBI)
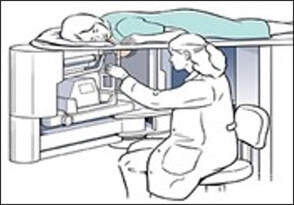
Using the X-ray / coordinate combination, the ABBI system uses an imaging table which displays pictures of the patient’s breast in snapshot form. A needle is then inserted into the selected area and a tissue core from 5 mm to 20 mm in diameter is removed. ABBI has been available since 1996, and marked a significant advance over the previous biopsy technique of wire localization
Using the X-ray / coordinate combination, the ABBI system uses an imaging table which displays pictures of the patient’s breast in snapshot form. A needle is then inserted into the selected area and a tissue core from 5 mm to 20 mm in diameter is removed. ABBI has been available since 1996, and marked a significant advance over the previous biopsy technique of wire localization
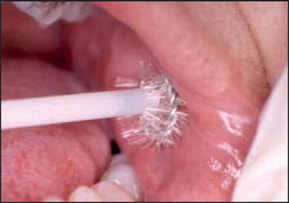
Brush biopsy
This technique is used for sampling the ureter as well as for detecting cancers in the mouth, bronchus, biliary tract and oesophagus. For tissue samples of the ureter for example, the method of retrograde brush biopsy cytology is used. In this method, a cystoscope enters through the urethra and the bladder in order for a guidewire to gain access to the ureter. A catheter is then passed over the guide wire and a contrast dye is instilled through it which enables positioning next to the lesion using fluoroscopy. After irrigation with saline solution, a nylon or steel brush is placed through the catheter and the lesion is rubbed with the brush. This is repeated several times using a new brush each time. When the brush is removed, the tip of the brush is saved and tissue from the lesion is removed from the brush tip. After the last brushing, the area is irrigated with saline and this is also sent for examination
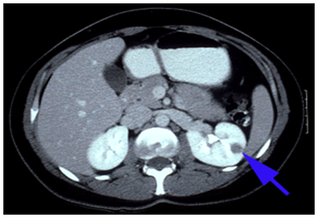
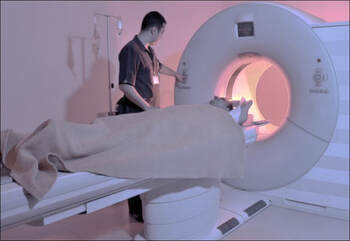
Computed tomography (CT)
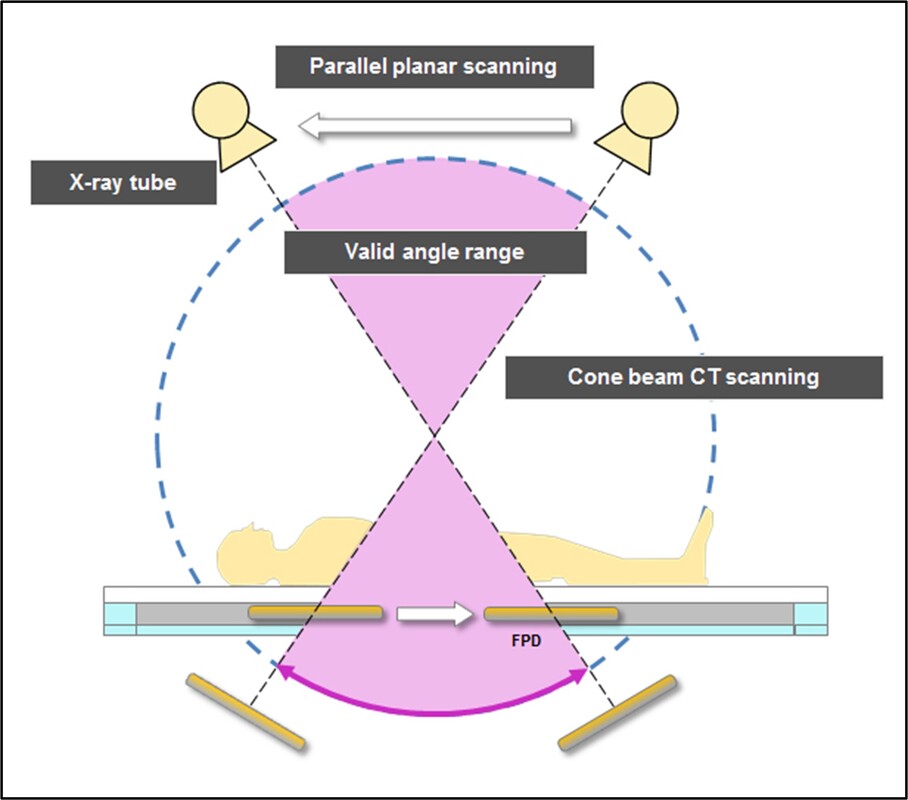
This is a diagnostic technique in which the combined use of a computer and X rays passed through the body at different angles produces clear cross-sectional images of the tissue under examination. CT scanning, also known as computed axial tomography (CAT scanning) is particularly useful for locating and imaging tumours and for facilitating needle biopsies
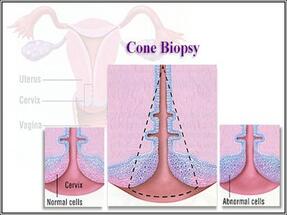
Cone biopsy
In this specialized technique, a small cone-shaped sample of tissue is removed from the inner surface of the cervix. Cone biopsies are used to diagnose cervical cancer in patients with either an abnormal cervical smear or punch biopsy
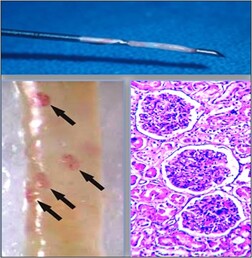
Core biopsy
These are performed in much the same way as a fine needle aspiration but with a larger bore needle. After a local anaesthetic, a small nick is made in the skin with a scalpel and the biopsy needle is injected through the nick and into the lump, removing a core of tissue. Core biopsies of kidney and palpable breast abnormalities may be performed with or without image guidance and usually provides a reliable histological diagnosis
Crosby-Kugler capsule
Capsule biopsies of jejunal mucosa for the investigation of malabsorption states can be assessed by stereomicroscopy where the architecture of the villous surface is clearly seen. Correct orientation of the mucosa will make interpretation easier
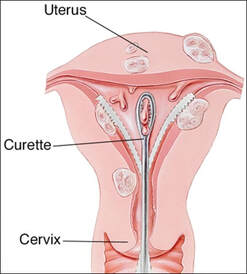
Curettings
Curetted specimens are awkward to orientate, but this is often only of importance with skin lesions. While there is little problem in identifying simple warts in these samples, differentiating between squamous cell carcinomas and other lesions may prove difficult. On the other hand, uterine curettings are usually obtained in sufficient quantity for adequate diagnosis
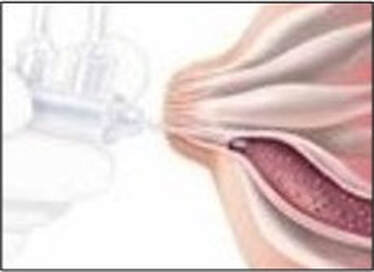
Ductal lavage
This test is carried out by inserting a tiny catheter into the nipple where a small amount of fluid is removed from the ductal cells after a saline flush
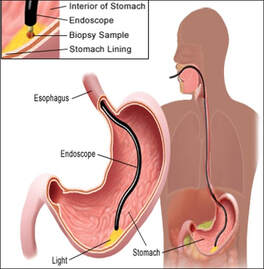
Endoscopic biopsy
This is a general term that refers to biopsy samples obtained by using endoscopes, special fibre optic instruments with miniature forceps at the end that can be passed into certain parts of the body. They are commonly used to take biopsy samples from areas such as the stomach (gastroscopy), the colon (colonoscopy or sigmoidoscopy), the bladder (cystoscopy), the lungs (bronchoscopy), the joints (arthroscopy) and the cervix (colposcopy). Laparoscopes are endoscopes for the examination of the abdominal cavity and are used in laparoscopic (keyhole) surgery for the removal of tissues such as the gall bladder
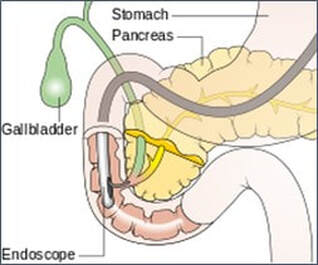
Endoscopic retrograde cholangiopancreatography (ERCP)
This is a technique that combines the use of endoscopy and fluoroscopy to diagnose and treat certain problems of the biliary or pancreatic ductal systems. Through the endoscope, the physician can see the inside of the stomach and duodenum, and inject radiographic contrast into the ducts in the biliary tree and pancreas so they can be seen on X-rays. ERCP is used primarily to diagnose and treat conditions of the bile ducts and main pancreatic duct, including gallstones, inflammatory strictures (scars), leaks (from trauma and surgery), and cancer. ERCP can be performed for diagnostic and therapeutic reasons, although the development of safer and relatively non-invasive investigations such as magnetic resonance cholangiopancreatography (MRCP) and endoscopic ultrasound has meant that ERCP is now rarely performed without therapeutic intent
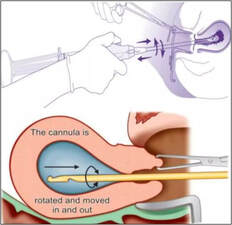
Evacuation
Surgical evacuation of the uterus for management of incomplete abortion, usually involves vacuum aspiration using a cannula
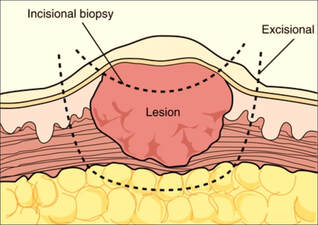
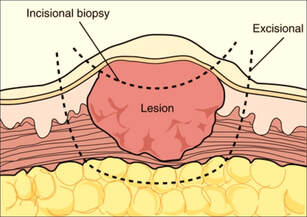
Excision biopsy
This is a surgical procedure performed on a lump that is small enough to be removed easily. Sometimes an entire tumour with a small rim (or margin) of normal tissue around it is removed. This technique is commonly used to remove relatively small and easily accessible masses. Surgeons should always mark the excisional biopsy margins with sutures or metal clips so that if removal is incomplete and further excision is needed the margin of previous excision can be properly located (compare with incisional biopsy)
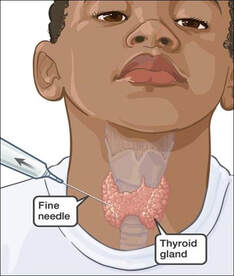
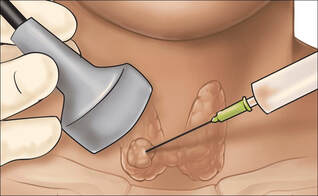
Fine needle aspiration (FNA)
This is the process of extracting cells from a lump such as that found in the breast by using a very thin, fine, hollow needle and syringe. This technique is used alone when the mass can be clearly palpated through the skin. For subtle or deep lesions the procedure is used along with X-ray or ultrasound guidance
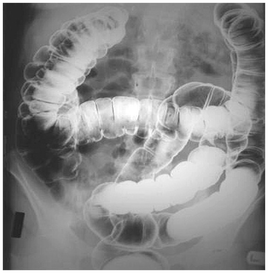
Fluoroscopy
This is a method for obtaining X-ray images and is used during many diagnostic and therapeutic procedures. Fluoroscopy is also used to observe the action of instruments during biopsy techniques. During this procedure, a transmitted X-ray beam strikes a fluorescent plate which is coupled to an image intensifier and monitor. Fluoroscopy is often used to observe the digestive tract after a barium enema
Frozen section
This is a rapid process that allows the pathologist to give the surgeon an immediate diagnosis while the patient is still in the operating room. In this procedure, a fresh tissue sample is removed and sent to the laboratory where it is frozen and cut in a cryostat. Microscopic slides are then prepared and examined by a pathologist who can determine whether the tissue is benign or malignant
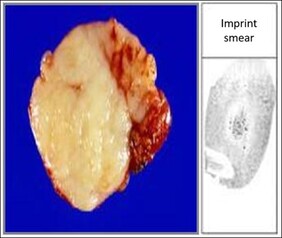
Imprint smear
This is a method whereby a smear is prepared by pressing the cut surface of a freshly dissected sample (such as lymph node) onto the surface of a microscope slide. This results in a thin layer of cells which can be stained and examined microscopically
Incision biopsy
This refers to a surgical technique used to sample a large, sometimes inaccessible mass which cannot be easily removed. The surgeon cuts into the mass and removes a sample which is then used to establish a definitive diagnosis of cancer before performing major surgery. Orientation of biopsy incisions is extremely important since ill-conceived incisions can unnecessarily open up additional tissue planes, necessitating more extensive ultimate surgical resections (compare with excisional biopsy)
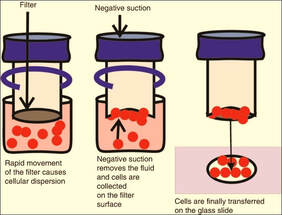
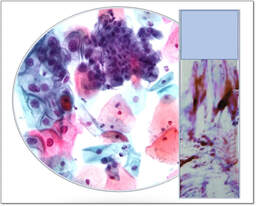
Liquid based cytology (LBC)
This is an automated alternative to the conventional cervical (Pap) smear. In the LBC (Thinprep) system, a sample from the cervix is collected using a brush type plastic device which is then detached into a vial of transport medium. In the laboratory, the tubes are vortex mixed and the suspension passed through a density gradient centrifugation process to remove mucus and blood cells. The cell pellet is then resuspended and a thin layer sample transferred to a microscope slide which can then be stained and examined microscopically
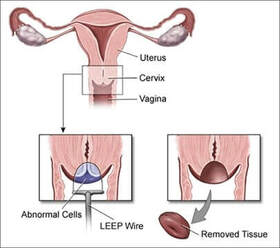
Loop electrosurgical excision procedure (LEEP)
This is one of the most commonly used approaches to treat high grade cervical dysplasia discovered on colposcopic examination. In the UK it is known as large loop excision of the transformation zone (LLETZ).The procedure has many advantages including low cost, high success rate, and ease of use. The procedure can be done in an office setting and usually only requires a local anaesthetic
Magnetic resonance imaging (MRI)
This is a non-invasive imaging technique which uses a strong magnet and radiofrequency waves to produce images of internal organs. The clinical role of MRI in cancer imaging includes determining the true edges of tumours prior to surgery, differentiating palpable masses from scar or dense tissue and detecting occult breast cancers mammographically and sonographically in patients with axillary nodal metastases. Though MRI has potential applications, it is not routinely used in breast cancer
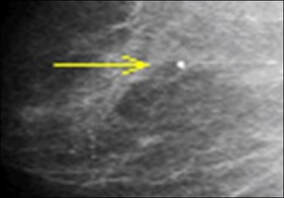
Mammography
The mammogram is an X-ray of the breast which is able to detect cancer at an early and curable stage. Mammography can identify breast cancers that are too small to be palpated by physical examination and can also detect the presence of localised lumps. Digital mammography is an emerging technique which uses computers and specially designed detectors to produce a digital image of the breast that can be displayed on high-resolution monitors. However, these tests cannot say whether the cancer is benign or malignant so a biopsy is needed to confirm the diagnosis. Vacuum-assisted breast biopsies are percutaneous procedures that rely on stereotactic mammography or ultrasound imaging. Stereotactic mammography involves using computers to pinpoint the exact location of a breast mass based on mammograms taken from two different angles. The computer coordinates will help the physician guide the needle to the correct area in the breast
Needle biopsy
In this technique, a large bore needle is inserted into a tissue mass to extract one or many cores of tissue. If a suspected tumour is inaccessible or located deep within the body, it may be necessary to use ultrasound imaging or a CT (computerised tomography) scanner in order to precisely position a needle through which a biopsy sample can be removed
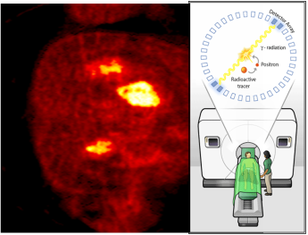
Positron emission tomography with fluorodeoxy- glucose (PET with FDG)
The PET technique involves the use of radioactive material in the diagnosis of cancer. In this method, the patient is injected with the radioactive substance fluorodeoxyglucose (FDG), a compound taken up by metabolically active tissues. As cancer cells are more metabolically active than normal tissues, the tumour tissue will take up relatively more radiolabelled substance. The patient is placed in a scanner to detect the radiation. This test may be particularly useful to determine the spread of cancer to other sites in the body. The main advantage of PET is that it can diagnose diseases even before the structural changes are visible. Since, it utilizes isotopes of basic biological elements like carbon, oxygen, and nitrogen, it reveals the disease status at a more cellular level than other types of imaging techniques
Punch biopsy
The instrument used is a hollow tube that is rotated into the skin until a core of tissue from the proper depth is obtained. This technique is used to obtain a deep core of skin and underlying tissue but is also used to examine the cervix following an abnormal cervical smear
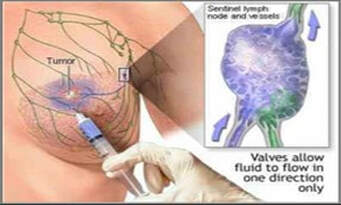
Sentinel node biopsy
This is used to check if an existing breast cancer has spread to lymph nodes. In the procedure, dye or radioactive material is injected into the region of a tumour which is then followed as it moves toward the lymph nodes. The first one it reaches is the sentinel which is then removed for histology
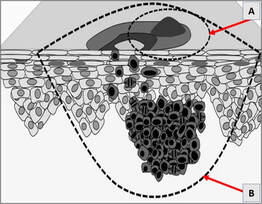
Shave and scoop biopsy
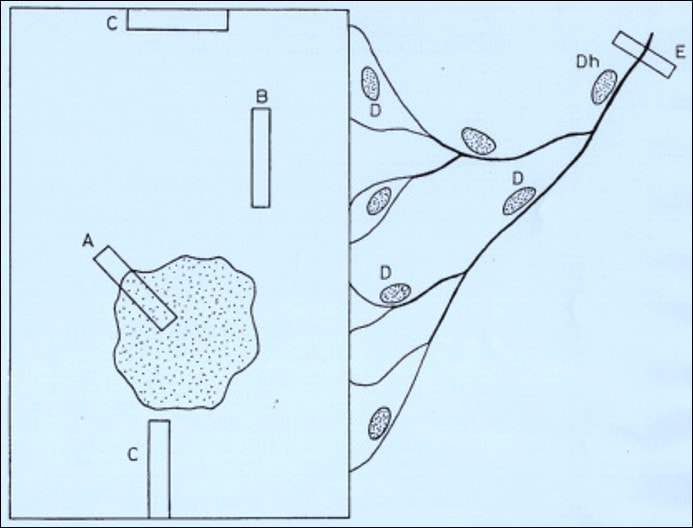
This is a method of obtaining a biopsy sample from a skin lesion. A scalpel is used to literally shave off the surface of a lesion and is commonly used to evaluate pigmented moles (suspected melanomas) or other tumours involving the skin (A). If the lesion is malignant then the scoop procedure will be necessary to remove the remaining portion (B)
Smears
In a smear, individual cells are removed using either a very thin needle or by scraping a surface with an instrument. The cells obtained are carefully smeared onto a glass slide, stained and examined microscopically. Cervical (Papanicolaou or Pap) smears are routinely used to examine cells from the surface of the cervix. In this procedure, the surface of the cervix is scraped with a spatula-shaped instrument to obtain a sample of cells
Stereotactic biopsy
This is a specialized radiology technique used to evaluate masses that are either too small to be felt directly through the skin or located in an inaccessible part of the body. It is most frequently used to evaluate breast masses where X-rays from two angles and a computer are used to locate the lump, and then a core needle is inserted into the breast for a tissue sample. With sophisticated radiological equipment, the suspicious mass is localized and a small incision made directly above. The radiologist or surgeon then uses a needle to extract several cores of tissue
Tomosynthesis
X-ray technology that offers high precision, multi-slice imaging at low exposure to visualize areas that are invisible by conventional radiography
Transurethral resection
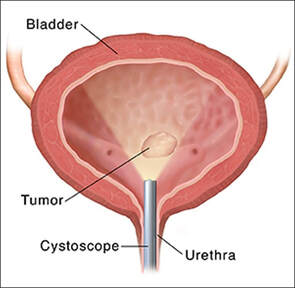
This involves surgical removal of the prostate or bladder lesions by means of an cystoscope inserted through the urethra, usually for the relief of prostatic obstruction or for treatment of bladder malignancies
Trephine biopsy
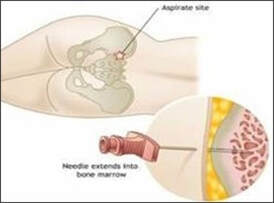
Bone marrow is located in the centre of many bones and is responsible for producing blood cells. The needle trephine removes a sample of the marrow which is used for the diagnosis and staging of cancers involving the blood cells which include lymphoma, leukaemia and metastatic disease
Ultrasound
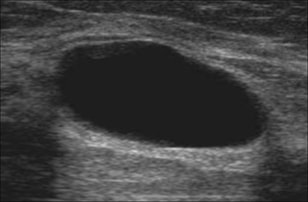
Ultrasound is used to help evaluate suspicious lesions in pre-menopausal women or evaluate breast abnormalities found on a mammogram. The ultrasound method comprises a small transducer which is able to slide along the skin, sending short bursts of high-frequency sound waves into the tissue. The sound waves will either bounce off the tissue or pass through it. The patterns of the sound waves that bounce off the tissue produce a picture of the mass from which it is possible to ascertain whether the mass is solid, semi-solid or a fluid-filled cyst
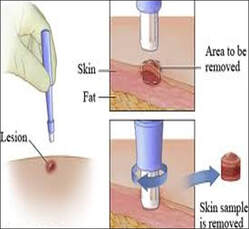
Vacuum-assisted biopsies
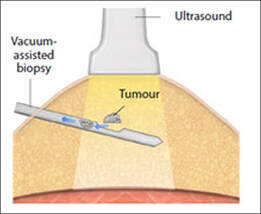
These are usually used for sampling breast tissue and are referred to by their brand names of Mammotome or MIBB (Minimally Invasive Breast Biopsy). The Mammotome procedure uses a tiny hollow needle probe, which is guided to the abnormal area by computer images projected on two screens (a stereotactic view) or by ultrasound. The needle is inserted into the breast tissue and uses a vacuum to withdraw the tissue, which is removed by a high-speed rotating cutter. The probe can obtain more than one sample, if necessary, without being withdrawn and reinserted. The physician simply repositions it and activates another rotation. A physician gets about 10 times as much breast tissue from the Mammotome procedure as from a core needle biopsy. The MIBB instrument resembles a radiolucent needle, making it easier to see on an X-ray. A specimen collection chamber is located below the tip with a vacuum unit that pulls the tissue into the chamber
Wire localization biopsy
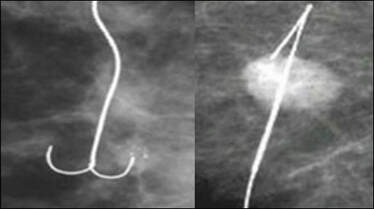
This is a specialized procedure used for small or non-palpable breast masses. Using X-ray or ultrasound guidance, the mass is located under anaesthetic and a thin wire is carefully inserted into the suspicious area. The surgeon uses the tip of the wire as a guide to locate and remove a sample for histology
Adverse effects of tissue sampling
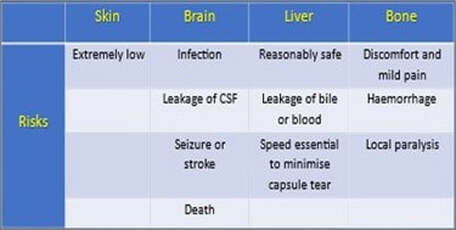
Irrespective of how careful biopsies are carried out, they are invasive techniques and as such carry elements of risk. Consequently, in order for patients to benefit from these procedures, there must be a real clinical indication for carrying them out. Risks with certain types of biopsy such as those from the skin are far less hazardous than with those of the liver for example. The risks of some of the more common biopsy sites are outlined below:
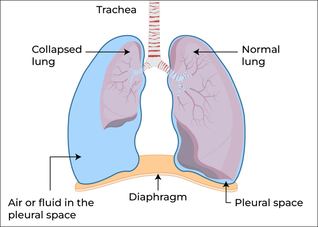
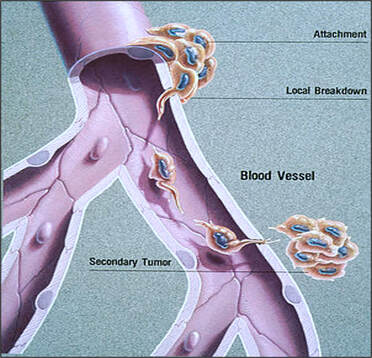
Specific risks of biopsy procedures include infection from a contaminated biopsy instrument or site, perforation of a tissue or organ, vascular problems such as bleeding from the biopsy site, dissemination of the tumour and induction of reactive changes which on subsequent biopsy may be misinterpreted. However, depending on the localization, other potential complications such as local paralysis, pneumothorax and sepsis may occur. With the stereotactic brain biopsy for example, the risks include stroke, seizure and even death.
Vascular damage Perforation of tissue or organ
Induction of reactive change Infection by contamination
Dissemination of tumour
Sampling of tissue inside the laboratory
The ability to accurately examine, describe and process gross specimens is one of the most important skills of the pathologist. Although this is rather obvious for the processing of entire biopsy specimens, the description of large surgical specimens provides a permanent record of all relevant information. This should include all the demographic and clinical information provided by the submitting physician, the observations made at the time of dissection and a description of samples taken. At the time of gross examination, resection margins of small biopsies are not generally marked with ink since delineation of surgical margins is not necessary. Simple specimens such as polyps with clear anatomic landmarks can be marked with ink at their surgical margins if required. With few exceptions, tissue samples arrive in the laboratory immersed in the fixative formalin (see the chapter on 'Formalin and tissue fixation').
Sampling of tissue inside the laboratory
The ability to accurately examine, describe and process gross specimens is one of the most important skills of the pathologist. Although this is rather obvious for the processing of entire biopsy specimens, the description of large surgical specimens provides a permanent record of all relevant information. This should include all the demographic and clinical information provided by the submitting physician, the observations made at the time of dissection and a description of samples taken. At the time of gross examination, resection margins of small biopsies are not generally marked with ink since delineation of surgical margins is not necessary. Simple specimens such as polyps with clear anatomic landmarks can be marked with ink at their surgical margins if required. With few exceptions, tissue samples arrive in the laboratory immersed in the fixative formalin (see the chapter on 'Formalin and tissue fixation').
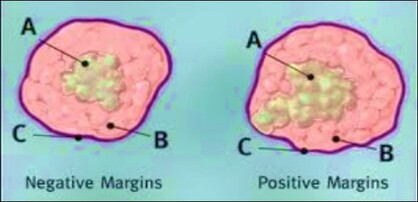
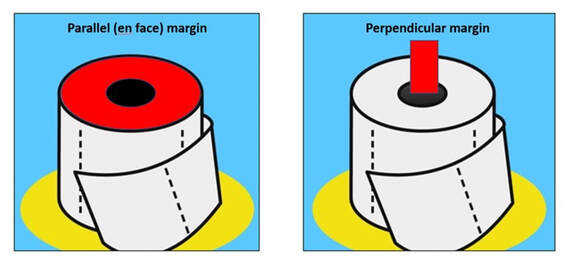
All specimens obtained from surgery with intent to cure should be inked at their surgical margins. In the laboratory, methods for assessing the specimen margins may either be parallel to the plane of resection (en face) or perpendicular to the plane of resection. The advantage of sampling a margin en face is that a large surface area can be examined as seen with the entire circumference of bowel resection margins. Although the margin can be called completely negative if no tumour is seen, the disadvantage is that the exact distance of the tumour to the margin cannot be determined.
Importantly, if any tumour is identified in an en face margin, the margin must be considered positive. The advantage of sampling a margin perpendicularly is the ability to determine precisely the distance of tumour from the margin. Thus, perpendicular margins are recommended when the tumour approaches closely (within 2.0 mm) to what otherwise would be considered a negative margin. The disadvantage is that very little of the margin is actually sampled. Margins are optimally inked and allowed to dry while the specimen is fresh, prior to preliminary dissection. Generally, rules for surgical resection margins are that all tissue samples with intent to cure should be inked. Both proximal and distal margins should be sampled wherever possible to include perpendicular to plane of resection or parallel to plane of resection.
If the tissue is received unfixed, frozen sections, imprint smears or samples for microbiological investigations may also be taken before the tissue is fixed in 10% formalin or other preferred fixative. At cut up, the pathologist can integrate the gross and microscopic appearances to arrive at a diagnosis. Surgical resections for tumour must be accurately described and measured with photographs and diagrams showing the sites of the selected tissue blocks. From these samples, the histopathologist will not only be able to diagnose the tumour type but will also be able to report on the extent of spread, the adequacy of resection and the presence of local precancerous lesions. Samples that include the edge of a tumour are usually more informative and although a single block is invariably sufficient, several blocks are usually taken. Sampling of blood vessels should also be performed to show the presence of vascular permeation. This should be performed in conjunction with the thorough sampling of nodes to show the extent of lymph node involvement and it is important that a standard sampling method is followed. Palpation of nodes is best avoided since it biases the sampling in favour of involved nodes. An alternative and better method is to slice the fat at regular intervals and process all nodes presenting at the cut surfaces.
Large format histology
In cancer pathology, the application of large format tissue processing is an established method for assessing characteristics such as intra-tumour heterogeneity, distribution and surgical margin involvement of large tumour samples. This method has proven to be cost effective and is able to meet the needs of the laboratory in the multidisciplinary approach to cancer diagnosis. Follow the links below for more information on methods of dissecting and processing large format tissue samples or you can check them out via the 'Publications' tab above.
Large format histology
In cancer pathology, the application of large format tissue processing is an established method for assessing characteristics such as intra-tumour heterogeneity, distribution and surgical margin involvement of large tumour samples. This method has proven to be cost effective and is able to meet the needs of the laboratory in the multidisciplinary approach to cancer diagnosis. Follow the links below for more information on methods of dissecting and processing large format tissue samples or you can check them out via the 'Publications' tab above.
Because antibiotic therapy is generally the first treatment of choice for specific infective processes, surgical resections are rarely performed unless they are found unexpectedly during surgery. Fresh samples of these tissues must be taken for microbiological studies before tissues are fixed. Culture is essential since some infections may only show sparse organisms that may not be detected by Gram stain or Ziehl Neelsen in paraffin sections. Resections and resection margins for chronic inflammatory bowel disease need to be sampled carefully because the lesions and granulomas may be sparse and patchy and precise location will therefore be required.
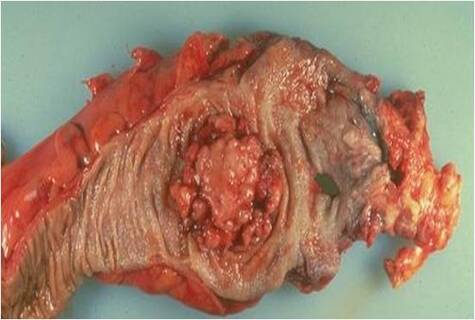
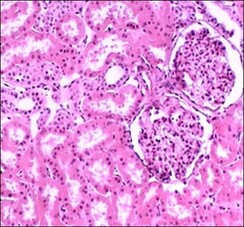
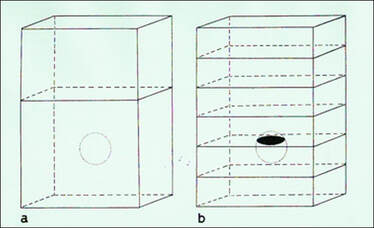
The difference between the amount of tissue sample taken and the object as a whole is known as sampling error and it may or may not be representative. The fact that changes in some diseases are localised rather than diffuse can cause diagnostic problems if sufficient slicing of the tissue is not performed (see figures below). By maintaining consistency, this can be overcome by taking what is regarded as an adequate number of blocks of larger pieces of tissue or resections. However, in small samples where only a single block is available, then it is equally important to examine deeper sections throughout the block to ensure that sampling errors are minimised.
For most diagnostic purposes, paraffin sections are cut at about 5 μm, although there are some uses for sections to be cut thicker or thinner. Thick sections will reduce sampling error but are virtually useless for diagnosis of most lesions because of superimposition. However, a 20 μm section can be a useful way to find pathogens such as bacteria or asbestos fibres provided the microscope is focused through all planes of the section during the examination. Frozen sections too, can be cut at 10 μm or greater and still provide a diagnosis particularly with staining methods associated with the nervous system. Although paraffin wax sections can be cut thinner than 5 μm, it is easier to use tissues that have been embedded in acrylic or epoxy resins since it provides greater support. Current applications include renal biopsies, lymph node biopsies and bone marrow trephines. The clarity provided by 1 μm plastic sections is due to the virtual elimination of superimposition of cells and to the considerable reduction in tissue shrinkage when compared with paraffin wax sections.
Diagnostic histopathology is an important requirement for both clinical practice and the management of patient care. The evidence provided of the many processes involved has shown that the interaction between the clinician and the laboratory is as vital as the ability of the pathologist to correctly interpret different abnormal states. Although microscopic examination of tissues is the main histological method of diagnosis, the technology of the modern era has changed all that. Storage of photographs as whole slide images has transformed the way that the processes of interpretation and diagnosis are now managed. The viewing experience of digital images to that of conventional microscopy is described at the end of the 'Staining' section.
Website updated 4th July 2024